 If you unravelled and stretched out the set of DNA molecules that reside in the nucleus of the tiniest human cell, they would extend 5 feet end-to-end. During cell division, however, these molecules tightly wind themselves up into microscopic packages called chromosomes. John Kyrk's website has a wonderful flash animation that shows the 3D structure of chromosomes: how the DNA molecule is wound around cylindrical proteins called histones, then further wound into a fibre of chromatin which is then finally wound up to form chromosomes. The information density this packaging achieves is staggering: 1.88 x 1021 bits (2 billion gigabytes) of information per cubic centimeter. Oh, and that includes error correction logic. No wonder DNA scaffolding is being looked at as the basis for a possible nanotech storage media.
The nucleus of most human cells contains 2 sets of chromosomes, 1 set given by each parent. Each set has 23 single chromosomes: 22 autosomes [common to both sexes] and an X or Y sex chromosome. (A female will have a pair of X chromosomes; a male will have one X and one Y.) (ref.)
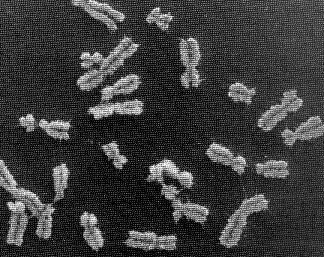
The picture at the start of this blog entry shows chromosome units that are in the process of duplicating themselves. The symmetric halves of the chromosome are called sister chromatids, both of which contain identical DNA molecules. Each chromatid (and the original chromosome that was split in two to form them) contain two end caps called telomeres and a centromere that forms its waist. (ref.)
If you unravelled and stretched out the set of DNA molecules that reside in the nucleus of the tiniest human cell, they would extend 5 feet end-to-end. During cell division, however, these molecules tightly wind themselves up into microscopic packages called chromosomes. John Kyrk's website has a wonderful flash animation that shows the 3D structure of chromosomes: how the DNA molecule is wound around cylindrical proteins called histones, then further wound into a fibre of chromatin which is then finally wound up to form chromosomes. The information density this packaging achieves is staggering: 1.88 x 1021 bits (2 billion gigabytes) of information per cubic centimeter. Oh, and that includes error correction logic. No wonder DNA scaffolding is being looked at as the basis for a possible nanotech storage media.
The nucleus of most human cells contains 2 sets of chromosomes, 1 set given by each parent. Each set has 23 single chromosomes: 22 autosomes [common to both sexes] and an X or Y sex chromosome. (A female will have a pair of X chromosomes; a male will have one X and one Y.) (ref.)
The picture at the start of this blog entry shows chromosome units that are in the process of duplicating themselves. The symmetric halves of the chromosome are called sister chromatids, both of which contain identical DNA molecules. Each chromatid (and the original chromosome that was split in two to form them) contain two end caps called telomeres and a centromere that forms its waist. (ref.)  It's amazizng how quickly DNA can wrap itself up into these packages: Fluorescence videomicroscopy and scanning force microscopy were used to follow, in real time, chromatin assembly on individual DNA molecules immersed in cell-free systems competent for physiological chromatin assembly. Within a few seconds, molecules are already compacted into a form exhibiting strong similarities to native chromatin fibers. In these extracts, the compaction rate is more than 100 times faster than expected from standard biochemical assays. (ref.)
Prior to a cell dividing, the chromosomes are replicated. The sister chromatids that result are initially held together by something called 'chromsome cohesion'. Chromosome cohesion is established during S phase (when the chromosomes are replicated) and is then dissolved completely in metaphase to allow sister chromatids to come apart. The dissolution of cohesion is highly regulated; human cell lines that have defects in the regulation of cohesion show the hallmarks of cancer cells. Furthermore, it has been suggested that the abnormal karyotypes that result in diseases such as Down syndrome are the result of the improper dissolution of chromosome cohesion (ref.)
It's amazizng how quickly DNA can wrap itself up into these packages: Fluorescence videomicroscopy and scanning force microscopy were used to follow, in real time, chromatin assembly on individual DNA molecules immersed in cell-free systems competent for physiological chromatin assembly. Within a few seconds, molecules are already compacted into a form exhibiting strong similarities to native chromatin fibers. In these extracts, the compaction rate is more than 100 times faster than expected from standard biochemical assays. (ref.)
Prior to a cell dividing, the chromosomes are replicated. The sister chromatids that result are initially held together by something called 'chromsome cohesion'. Chromosome cohesion is established during S phase (when the chromosomes are replicated) and is then dissolved completely in metaphase to allow sister chromatids to come apart. The dissolution of cohesion is highly regulated; human cell lines that have defects in the regulation of cohesion show the hallmarks of cancer cells. Furthermore, it has been suggested that the abnormal karyotypes that result in diseases such as Down syndrome are the result of the improper dissolution of chromosome cohesion (ref.)Cohesin sites (red ovals) are concentrated at the centromere/pericentric region (where the two chromatids are “pinched”), but also occur along the arms of the chromatids. MIT recently discovered that a protein called MEI-S332 regulates the chromosome cohesion, releasing the chromosomes from each other by adding a phosphate to the binding point. These findings are particularly significant given that researchers have found that levels of MEI-S332 are higher than normal in 90% of all breast cancers. According to Clarke, this might mean that when there’s too much of the protein, the chromosomes don’t separate properly, or it might mean that the MEI-S332 gene is mutated on the chromosomes. (ref.) During cell division (mitosis), the two sister chromatids of each of the 46 chromosomes are pulled apart. Protein filaments called microtubules attach the centromeres of the sister chromatids of the chromosomes to the cytoskeleton on opposite sides of the cell. The cytoskeleton is located along the inside of the cell membrane and acts like a nano-scale motor, pulling on the microtubules to line up the chromosomes in the middle of the cell and then literally yanking the sister chromatids apart. There's a great animation by HybridMedicalAnimation of this available here. There's also a fascinating movie of a plant cell undergoing mitosis (more here). It took me a while to appreciate that the structural packaging of DNA differs in bacteria, plants and animals. More:
'From a single double helix to paired double helices and back' The detailed mechanics involved in aligning the chromosomes down the middle of the cell is covered in this article.
No comments:
Post a Comment